Cell-Matrix Receptors[Edit]
Interaction between cell-matrix receptors and their respective ligands are often the initial step in the formation of a cell-matrix adhesion. Several types of cell-matrix receptors have been identified, each interacting with a specific type of ligand. Attachment of these various ECM based ligands or molecules to the exterior portion of the adhesion receptor causes their structural rearrangement [1]. This may be induced by a specific chemical property or change, (reviewed in [2]), a change in topography [3, 4, 5] (reviewed in [6, 7, 8]), or even the rigidity of the ECM components [9, 10].
 Figure 1. Integrins as adhesion receptor in focal adhesion (FA): Integrins upon binding to ECM ligand gets activated and undergoes conformational changes. This leads to a series of events including i) recruitment of binding proteins to the site leading to actin linking, ii) activation of signaling molecules leading to integrin clustering and iii) actomyosin contractions leading to adhesion strengthening. All these result in iv) formation of new actin filaments by actin polymerization modules.Cell-matrix adhesion receptors are grouped according to the ligand that is bound. In some cases, adhesion receptors bind more than one type of ligand (e.g. the integrin family) and different receptor groups may cooperate with each other to bind their ligands (reviewed in [11, 12]).Ligand-receptor binding is followed by the rapid association of other proteins to the intracellular portion of the receptor; this reinforcement of the adhesion domain is controlled by adhesion receptor mobility in the membrane [13]. Such change in forces can affect mechanosensory molecules to activate intracellular signal transduction cascades (e.g. the Rho family of GTPases) and mechanotransduction events that mediate a number of diverse processes such as cell proliferation, fate, migration, shape and polarization [14, 10] (reviewed in [15, 6, 16, 17, 18]).
Figure 1. Integrins as adhesion receptor in focal adhesion (FA): Integrins upon binding to ECM ligand gets activated and undergoes conformational changes. This leads to a series of events including i) recruitment of binding proteins to the site leading to actin linking, ii) activation of signaling molecules leading to integrin clustering and iii) actomyosin contractions leading to adhesion strengthening. All these result in iv) formation of new actin filaments by actin polymerization modules.Cell-matrix adhesion receptors are grouped according to the ligand that is bound. In some cases, adhesion receptors bind more than one type of ligand (e.g. the integrin family) and different receptor groups may cooperate with each other to bind their ligands (reviewed in [11, 12]).Ligand-receptor binding is followed by the rapid association of other proteins to the intracellular portion of the receptor; this reinforcement of the adhesion domain is controlled by adhesion receptor mobility in the membrane [13]. Such change in forces can affect mechanosensory molecules to activate intracellular signal transduction cascades (e.g. the Rho family of GTPases) and mechanotransduction events that mediate a number of diverse processes such as cell proliferation, fate, migration, shape and polarization [14, 10] (reviewed in [15, 6, 16, 17, 18]).
Various types of cell-matrix receptors exist. These include:
* Fibronectin receptors, which include the most common types of adhesion receptors, the integrin family and the syndecan family of transmembrane proteoglycans.
* Collagen receptors, which includes integrins, receptor tyrosine kinases (e.g. discoidin domain receptors [DDRs]), glycoproteins (e.g. GPVI) and immunoglobulin-like proteins aka IgCAMs (e.g. LAIR-1).
* Laminin receptors, which include integrins, the dystrophin glycoprotein complex (DGC), the 67 kDa laminin receptor (67LR), and two glycoproteins belonging to the immunoglobulin superfamily, Lutheran (Lu) and basal cell adhesion molecule (B-CAM).
* Hyaluronan receptors (aka hyaladherins) Proteins that bind to hyaluronan, a large polysaccharide, have immunoglobulin-like domains and include members of the CD44 family and CD168.
CAMs have many distinct domains that allow them to mediate cell-cell contacts by binding to specific partner proteins; when these interactions occur between apposed cells they are described as either homophilic (i.e. binding to the same kind of CAM molecule) or heterophilic (binding to a different kind of CAM molecule). Furthermore, CAMs can mediate interactions between cells of the same type (aka homotypic adhesion) or between different cell types (aka heterotypic adhesion).
CAMs are grouped into four main families:
* Cadherins: Mediate primarily homophilic interactions at cell-cell adhesions.
* Immunoglobulin superfamily (Ig) CAMs: Mediate homophilic interactions at cell-cell adhesions.
* Integrins: Some integrin types mediate heterophilic cell-cell interactions.
* Selectins: Mediate heterophilic interactions at cell-cell adhesions.
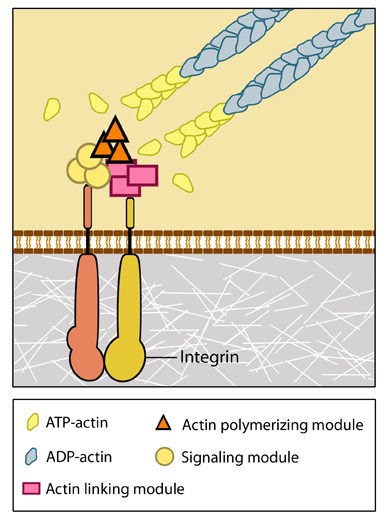 Figure 1. Integrins as adhesion receptor in focal adhesion (FA): Integrins upon binding to ECM ligand gets activated and undergoes conformational changes. This leads to a series of events including i) recruitment of binding proteins to the site leading to actin linking, ii) activation of signaling molecules leading to integrin clustering and iii) actomyosin contractions leading to adhesion strengthening. All these result in iv) formation of new actin filaments by actin polymerization modules.Cell-matrix adhesion receptors are grouped according to the ligand that is bound. In some cases, adhesion receptors bind more than one type of ligand (e.g. the integrin family) and different receptor groups may cooperate with each other to bind their ligands (reviewed in [11, 12]).
Figure 1. Integrins as adhesion receptor in focal adhesion (FA): Integrins upon binding to ECM ligand gets activated and undergoes conformational changes. This leads to a series of events including i) recruitment of binding proteins to the site leading to actin linking, ii) activation of signaling molecules leading to integrin clustering and iii) actomyosin contractions leading to adhesion strengthening. All these result in iv) formation of new actin filaments by actin polymerization modules.Cell-matrix adhesion receptors are grouped according to the ligand that is bound. In some cases, adhesion receptors bind more than one type of ligand (e.g. the integrin family) and different receptor groups may cooperate with each other to bind their ligands (reviewed in [11, 12]).Various types of cell-matrix receptors exist. These include:
* Fibronectin receptors, which include the most common types of adhesion receptors, the integrin family and the syndecan family of transmembrane proteoglycans.
* Collagen receptors, which includes integrins, receptor tyrosine kinases (e.g. discoidin domain receptors [DDRs]), glycoproteins (e.g. GPVI) and immunoglobulin-like proteins aka IgCAMs (e.g. LAIR-1).
* Laminin receptors, which include integrins, the dystrophin glycoprotein complex (DGC), the 67 kDa laminin receptor (67LR), and two glycoproteins belonging to the immunoglobulin superfamily, Lutheran (Lu) and basal cell adhesion molecule (B-CAM).
* Hyaluronan receptors (aka hyaladherins) Proteins that bind to hyaluronan, a large polysaccharide, have immunoglobulin-like domains and include members of the CD44 family and CD168.
Cell-Adhesion Molecules
Anchoring junctions are multiprotein complexes. Crucial to the formation of these junctions are cell adhesion molecules (CAMs).CAMs have many distinct domains that allow them to mediate cell-cell contacts by binding to specific partner proteins; when these interactions occur between apposed cells they are described as either homophilic (i.e. binding to the same kind of CAM molecule) or heterophilic (binding to a different kind of CAM molecule). Furthermore, CAMs can mediate interactions between cells of the same type (aka homotypic adhesion) or between different cell types (aka heterotypic adhesion).
CAMs are grouped into four main families:
* Cadherins: Mediate primarily homophilic interactions at cell-cell adhesions.
* Immunoglobulin superfamily (Ig) CAMs: Mediate homophilic interactions at cell-cell adhesions.
* Integrins: Some integrin types mediate heterophilic cell-cell interactions.
* Selectins: Mediate heterophilic interactions at cell-cell adhesions.